RHESSI Imaging Software
The Fourier-transform imaging method used by RHESSI is based on nine rotating modulation collimators (a pair of widely separated grids in front of a detector) that time-modulate the incident flux as the spacecraft rotates. During a solar flare, the counts recorded by the detectors are modulated in patterns that depend on the location, size, and intensity of the source. Different Fourier components of the angular distribution of the source are measured at different rotation angles and with grids of different pitches. The image reconstruction software attempts to determine the spatial photon distribution by analyzing these modulated time profiles and incorporating the knowledge of the instantaneous spacecraft pointing and roll position. Please see the RHESSI Imaging Concept for a detailed explanation of how RHESSI images. Another good explanation with some software examples is provided in the Introduction to RHESSI X-ray Imaging.
There are eleven image reconstruction algorithms, as well as two obsolete algorithms, all listed to the left. Each algorithm has a different method for solving this inverse Fourier-transform problem. There is no single 'best' algorithm; the best algorithm to use in a particular case depends on the source morphology and intensity, and your scientific goals. See the Image Algorithm Summary for a description of each algorithm and its strengths and weaknesses. Often, it helps to use the 'back projection' algorithm to generate an initial estimate of the image.
The units of the reconstructed images are photons cm-2 sec-1 arcsec-2.
RHESSI Image Software Basics
The RHESSI image software can be used from the IDL command line or the RHESSI GUI. The Introduction to RHESSI X-ray Imaging (referenced above, starting at slide 24) contains a good description of the Image GUI as well as some command line examples. Another useful guide to the Image GUI is provided in RHESSI Imaging - First Steps.
The image reconstruction software starts with an hsi_image object. By setting parameters in the image object, you control the image reconstruction. The image object produces either a single image, or an image cube depending on whether you set a single time and energy bin, or multiple times and/or energy bins (currently those are the only two dimensions allowed).
The image control and info parameters are listed in the Image Object Parameter Table. In addition to the general image parameters, each algorithm has a set of algorithm-specific parameters you can see by clicking on the name of the algorithm in the table header. The control parameters most commonly set in the image object by users are as follows (control parameter name in parentheses). Look in the table for more description of each parameter.
- time interval(s) (im_time_interval),
- energy band(s) (im_energy_binning),
- image algorithm (image_algorithm),
- subcollimator selection (det_index_mask),
- size of each pixel in image in arcsec (pixel_size),
- number of pixels in image in x and y (image_dim), and
- position of source on Sun (xyoffset) - only needed if position in flare catalog is missing or wrong.
To make an image from the command line, you create an image object, set your options, and request the data. As with all the RHESSI objects, all of the parameters are set to default values (shown in the parameter table, or by typing print,o->get(/xxx)) and you need to set only the values that you want to change. The only parameter for which there is no valid default is the time interval - you must set the im_time_interval parameter. For example,
o = hsi_image()
o -> set, im_time_interval= ['31-Oct-2010 01:15:50.000', '31-Oct-2010 01:15:54.000']
o -> set, image_algorithm= 'clean'
image = o -> getdata()
returns the image array for the specified time interval using the CLEAN algorithm.
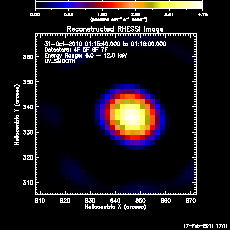
Here's an example in which we set all of the control parameters listed above and plot the image (shown on right):
o = hsi_image()
o -> set, im_time_interval= ['31-Oct-2010 01:15:40.000', $
'31-Oct-2010 01:16:00.000']
o -> set, im_energy_binning= [6., 12.]
o -> set, image_algorithm= 'UV_Smooth'
o -> set, det_index_mask= [0, 0, 0, 1, 1, 1, 1, 0, 0] ; use dets 4,5,6,7
o -> set, pixel_size= [2., 2.]
o -> set, image_dim= [32, 32]
o -> set, xyoffset= [840., 337.]
o -> plotman
Using Different Algorithms
The default image algorithm in the image object is Back Projection. To select a different algorithm:
o->set, image_algorithm='xxx'
There is more information on each algorithm in the tabs on the left, or in the Image Algorithm Summary.
In addition to the general image object parameters, each algorithm has a set of algorithm-specific parameters. The Image Object Parameter Table shows all of the general and algorithm-specific parameters; there are also separate tables for each algorithm showing only the parameters specific to that algorithm.
The parameters for a specific algorithm are not available in an image object until that algorithm has been selected. In the following example, we make a new image object, select the Clean algorithm, and try to retrieve the value of a Pixon control parameter, pixon_sizes. The value is -1 indicating it is not available.
o=hsi_image()
o->set,image_algorithm='clean'
print,o->get(/pixon_sizes)
-1
Now we set the image algorithm to Pixon, and retrieve the values for the pixon_sizes parameter.
o->set,image_algorithm='pixon'
print,o->get(/pixon_sizes)
16.5000 8.00000 5.65685 4.00000 2.82843 2.00000 1.41421 1.00000
Here, we set the image algorithm back to Clean. Because the Pixon algorithm had previously been selected in this instance of o, our image object, we can still retrieve the value for the pixon_sizes parameter:
o->set,image_algorithm='clean'
print,o->get(/pixon_sizes)
16.5000 8.00000 5.65685 4.00000 2.82843 2.00000 1.41421 1.00000
Grid Selection in Making Images
The selection of subcollimators and energy band used in constructing the image is important. Refer to the Grid Parameter Table for the FWHM resolution of each subcollimator and keep the following in mind:
- Subcollimators with a FWHM less than the smallest source size add no information, only noise.
- Subcollimators with a FWHM less than twice the pixel size will introduce alias features into the image.
- Subcollimators with a FWHM greater than the source extent only add integrated flux.
- Subcollimators that are transparent at the energy band selected only add integrated flux.
- Using too wide an energy band may result in errors because the instrument response used to compute the image is computed for the mean energy.
Using RHESSI Image FITS Files
The image(s) generated by the RHESSI imaging software can be stored in a FITS file along with all of the control and info parameters used to create it.
To write an image FITS file, use the fitswrite method. You can set the output file name by setting the im_out_fits_filename parameter, or if you don't want the parameter to persist, use the this_out_filename keyword on the call to fitswrite. For example:
o -> fitswrite, im_out_fits_filename='xxx.fits' ;sets and remembers im_out_fits_filename parameter
o -> fitswrite, this_out_filename='xxx.fits' ; xxx.fits filename is used just for this call
o -> fitswrite ; if im_out_fits_filename not set, prompts you for filename
To use the FITS file, you have several options.
- You can read the FITS file into an image object by setting the im_input_fits parameter. In the Image GUI, this is accomplished by setting 'Select Input' to 'Image FITS File', and then browsing for your FITS file. At the command line, you could read in your fits file as follows:
o->set, im_input_fits='xxx.fits'
data = o->getdata()
This sets all of the control and info parameters to their values when the FITS file was written and retrieves the image. As long as im_input_fits is set to 'xxx.fits', everything retrieved by calls to get and getdata is information that was stored in the file. You may not change any control parameters or do any reprocessing. To convert back to 'raw' mode (i.e. using the Level-0 raw files as input, and being able to change control parameters) set im_imput_fits to a blank string:
o->set, im_input_fits = ''
Note that all of the control parameters that were set when you read the FITS file are still set. However, now you are allowed to change control parameters, and a getdata will reprocess the images from the Level-0 data files. If you want to reinitialize the control parameters to default values, destroy the object and start with a fresh object. - Use the hsi_image_fitsread routine to return the image array(s) and/or the control or info parameters, or return an hsi_image object reconstructed from the FITS file. See the routine header for all of the options. A simple example:
image = hsi_image_fitsread(fitsfile='xxx.fits') - Use the hsi_fits2map routine to return a map structure (or an array of map structures for an image cube) from the FITS file. For example,
hsi_fits2map, 'xxx.fits', map
RHESSI Image Cubes
If you specify more than one time interval and/or energy band in the image object, then you will create an image cube. For example, the following settings will make an image cube for three time intervals and two energy bins:
o -> set, im_time_interval='20-feb-02 11:06:'+ [['00','04'], ['08','12'], ['16','20'] ]
o -> set, im_energy_binning = [[3,6], [6.,12]]
See How to Work with the Image Object for a full description of using the image object with image cubes, including
- alternate ways of setting the multiple time or energy bins,
- showing a movie,
- displaying all panels at once,
- choosing which images to show in a PLOTMAN session, and
- retrieving images and info parameters for all or particular images in the cube.
Data Stacking
All of the image algorithms can take advantage of data stacking. Data stacking involves combining data on the basis of spacecraft roll angle and aspect phase to make data from multiple rotations equivalent to that from a single rotation. The advantages are improved statistics, faster processing, and the possibility of imaging over longer time intervals.
Data stacking (also called phase stacking) is enabled in the Image GUI by enabling the 'Use Phase Stacker' button. At the command line, it's enabled by
o -> set, /use_phz_stacker
When you make an image with a visibility-based algorithm, the phase stacker is automatically enabled, since it is required.
Visibilities
Visibilities are complex numbers with amplitude and phase that are the result of fitting a sinusoid to the stacked profiles (so data stacking is required for visibility-based algorithms). They represent a compact, noise-free transformation of the input data.
Some of the image algorithms are based on visibilities. The visibility-based algorithms are MEM_NJIT, MEM_GE, VIS_FWDFIT, UV_SMOOTH, VIS_CS, and VIS_WV.
As of October 2012, you can also make regularized electron and photon visibilities and images. This is controlled in the Image GUI through the 'Visibility Type' pulldown menu. The options are 'photon' (the type of visibility available before Oct 2012), 'regularized photon', and 'regularized electron'. These options are not available until you select one of the visibiity-based image algorithms. From the command line, you can set the vis_type parameter to one of these values: 'photon', 'regularized photon', or 'regularized electron' or the shorthand names 'pho, 'reg pho' or 'reg el', e.g.
o->set, vis_type='reg el'
There are energy binning requirements you must meet when using regularized photon or electron visibilities.
See Using Visibility Objects in the RHESSI Software for details on using visibility objects and the visibility-based image algorithms, as well as more information about regularized visibilities.
See RHESSI Visibilities for more information about visibility FITS files and the Visibility FITS File Archive.
For more explanation of visibilities, see
- Measurement and Interpretation of X-ray Visibilities with RHESSI - Hurford, Schmahl, Schwartz, SPD 2005 Poster
- RHESSI Visibilities Nugget #39 - Schmahl and Hurford, 2006
- Wiki Article on RHESSI Visibilities - Christe, 2010
- Choosing the Number of Roll Bins for RHESSI Visibilities - Hurford, 2009
- User's Guide to RHESSI Visibilities - Hurford, 2006
- Optical Interferometry in Astronomy - John Monnier, 2003
Using Image Modulation Profiles or Visibility Profiles to Evaluate Image Quality
One good way to evaluate the reliability of a reconstructed image is to compare the observed count rate profile to the expected modulation profiles that would have produced the generated image. Another method is to compare the observed visibility amplitutudes and errors to the visibilities computed by the image.
See Image Modulation Profiles and Visibility Comparison Plots for details.



